Monday, November 4, 2013
Lab Skills You Stopped Being Proud Of
Wednesday, May 1, 2013
The Development of mNeonGreen
This week our most recent publication, “A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum” will be published in Nature Methods. It has already been viewable online for some time now, here is a link. We believe this new protein possesses a great deal of potential to advance the imaging fields through enhanced fluorescent microscopy. mNeonGreen enables numerous super resolution imaging techniques and allows for greater clarity and insight into one’s research. As a result of this we are taking a new approach at Allele for distribution of this protein, and here we will describe the history of the protein and some of the factors that led us down this path.
mNeonGreen was developed by Dr. Nathan Shaner at Allele Biotechnology and the Scintillon Institute through the directed evolution of a yellow fluorescent protein we offer called LanYFP. LanYFP is a super bright yellow fluorescent protein derived from the Lancelet fish species, characterized by its very high quantum yield, however, in its native state LanYFP is tetrameric. Dr. Shaner was able to monomerize the protein and enhance a number of beneficial properties such as photostability and maturation time. The result is a protein that performs very well in a number of applications, but is also backwards compatible with and equipment for GFP imaging.
Upon publication there was a question of how distribution should be structured. How would we make this protein available to researchers in a simple manner was a very difficult challenge. We also relied heavily on Dr. Shaner’s knowledge and experience in these matters, as he related his experiences to us from his time in Roger Tsien’s lab at UCSD. When the mFruits was published their lab was inundated with requests. The average waiting period was 3 months to receive a protein and they required a dedicated research technician to handle this process. Eventually the mFruits from the Tsien lab were almost exclusively offered through Clontech. Thus we decided that Allele Biotechnology would handle the protein distribution and take a commercial approach to drastically decrease the turnaround time. The next challenge we faced was how to charge for this protein. Due to the cost of developing this protein, which was fully funded by Allele, there is a necessity to recoup our investment and ideally justify further development of research tools, but we also understand the budget constraints every lab now faces. From this line of thinking we conceived our group licensing model; we wanted to limit the charge to $100 per lab. The way this is fiscally justifiable is having every lab in a department or site license the protein at this charge, including access to all related plasmids made by us as well as those generated by other licensed users (Click here for our licensing page). The benefit we see to this is that the protein is licensed for full use at a low cost, and collaboration amongst ones colleagues is not only permissible, it’s encouraged. We saw this as a win-win situation. We would recoup our cost and invest in further fluorescent protein research, and our protein costs would not be a barrier to research and innovation.
The granting of a license to use but not distribute material is not unique to commercial sources. Although academic material transfer agreements typically contain specific language forbidding distribution of received material beyond the recipient laboratory, some researchers choose to disregard these provisions. Unfortunately through this action they are disrespecting the intellectual property rights of the original researchers as well as violating the terms of the legal contract they signed in order to receive the material. We believe most researchers choose to respect the great deal of effort that goes into the creation of research tools for biology and do not distribute any material received from other labs without their express permission. However for a company that funds its own basic research our focus is often on the former example rather than the latter. We believe that this focus artificially drives up the costs of licensing a fluorescent protein and obtaining the plasmid, thus we have chosen to believe researchers will respect our intellectual property as long as we are reasonable in our distribution which is something we have truly striven for.
Additionally we believe the broad-range usage of a superior, new generation FP is an opportunity to advocate newer technologies that can be enabled by mNeonGreen, together with a number of Allele’s other fluorescent proteins (such as the photoconvertible mClavGR2, and mMaple). These new imaging technologies are called super resolution imaging (MRI). They provide researchers with a much finer resolution of cellular structures, protein molecule localizations, and protein-protein interaction information. We have started the construction of a dedicated webpage to provide early adopters with practical and simple guidance, click here to visit our super resolution imaging portal.
Friday, July 30, 2010
Allele’s pallet of the super star fluorescent proteins
http://allelebiotech.com/blogs/2010/07/alleles-pallet-of-the-super-star-fluorescent-proteins/
- The brightest cyan, green fluorescent proteins, and the brightest ever FP in LanYFP!
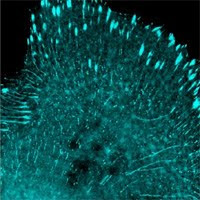
- Cells infected with lentivirus carrying mWasabi. Lentivirus carrying LanYFP will make most cells much more brighter than this.
- The LanFPs express well in bacteria.
- The FPs fold so strongly that they fluorescence even in SDS-PAGE.
- FPs in SDS PAGE–a closer look
- FPs in gel cassette over UV lights
- FPs in gel cassette under blue LED
Friday, March 19, 2010
Fluorescent Protein-Based Assay Development
Overview:
Originally cloned from the jellyfish Aequorea victoria and subsequently from many other marine organisms, fluorescent proteins (FPs) spanning the entire visual spectrum have become some of the most widely used genetically encoded tags. Unlike traditional labeling methods, FPs may be used to specifically label virtually any protein of interest in a living cell with minimal perturbation to its endogenous function. Genes encoding FPs alone or as fusions to a protein of interest may be introduced to cells by a number of different methods, including simple plasmid transfection or viral transduction. Once expressed, FPs are easily detected with standard fluorescence microscopy equipment.
Factors that should be taken into account when designing an FP-based imaging experiment include the desired wavelength(s) for detection, the pH environment of the tagged protein, the total required imaging time, and the expression level or dynamic range required for detection of promoter activity or tagged protein. Individual FPs currently available to the research community vary considerably in their photostability, pH sensitivity, and overall brightness, and so FPs must be chosen with care to maximize the likelihood of success in a particular experimental context.
FPs as fusion tags:
Use of FPs as fusion tags allows visualization of the dynamic localization of the tagged protein in living cells. For such applications, the cDNA of a protein of interest is attached in-frame to the coding sequence for the desired FP, and both are put under the control of a promoter appropriate to the experimental context (typically CMV for high-level expression, though other promoters may be desirable if overexpression of your protein of interest is suspected of producing artifacts). The most basic uses for fluorescent protein fusions include tracking of specific organelles (fusions to short organelle targeting signals) or cytoskeletal structures (fusions to actin or tubulin, for example). More advanced uses include tracking receptors or exported proteins. In most cases, it is critical that the FP used for fusion tagging be fully monomeric, as any interaction between fusion tags is likely to produce artifacts, some of which may be hard to recognize in the absence of other controls. While in most cases FP fusions do not interfere with normal protein function, whenever possible, FP fusion proteins should be validated by immunostaining the corresponding endogenous protein in non-transfected cells and verifying similar patterns of localization.
FPs as expression reporters:
FPs are highly useful as quantitative expression reporters. By driving the expression of an FP gene by a specific promoter of interest, it is possible to produce an optical readout of promoter activity. Use of the brightest possible FP ensures the best dynamic range for such an experiment. Because dynamic localization is not generally an issue for expression reporter applications, it is possible to use non-monomeric FPs for this purpose, opening up additional possibilities for multiple wavelength imaging. In order to obtain more reliable quantitative data and to correct for likely variations between individual cells in expression reporter experiments, the use of two spectrally distinct (e.g. green and red) FPs is advisable. By driving expression of one FP with a constitutive promoter and a second FP with the promoter of interest, the ratio of the two signals provides a quantitative readout of relative activity. Averaged over many cells, this technique should provide statistical power necessary for quality expression level experiments. Because FPs normally have a very slow turnover rate in mammalian cells, it may be desirable to add a degradation tag to your FP to enhance temporal resolution when measuring highly dynamic promoter activity.
Overview:
Originally cloned from the jellyfish Aequorea victoria and subsequently from many other marine organisms, fluorescent proteins (FPs) spanning the entire visual spectrum have become some of the most widely used genetically encoded tags. Unlike traditional labeling methods, FPs may be used to specifically label virtually any protein of interest in a living cell with minimal perturbation to its endogenous function. Genes encoding FPs alone or as fusions to a protein of interest may be introduced to cells by a number of different methods, including simple plasmid transfection or viral transduction. Once expressed, FPs are easily detected with standard fluorescence microscopy equipment.
Factors that should be taken into account when designing an FP-based imaging experiment include the desired wavelength(s) for detection, the pH environment of the tagged protein, the total required imaging time, and the expression level or dynamic range required for detection of promoter activity or tagged protein. Individual FPs currently available to the research community vary considerably in their photostability, pH sensitivity, and overall brightness, and so FPs must be chosen with care to maximize the likelihood of success in a particular experimental context.
FPs as fusion tags:
Use of FPs as fusion tags allows visualization of the dynamic localization of the tagged protein in living cells. For such applications, the cDNA of a protein of interest is attached in-frame to the coding sequence for the desired FP, and both are put under the control of a promoter appropriate to the experimental context (typically CMV for high-level expression, though other promoters may be desirable if overexpression of your protein of interest is suspected of producing artifacts). The most basic uses for fluorescent protein fusions include tracking of specific organelles (fusions to short organelle targeting signals) or cytoskeletal structures (fusions to actin or tubulin, for example). More advanced uses include tracking receptors or exported proteins. In most cases, it is critical that the FP used for fusion tagging be fully monomeric, as any interaction between fusion tags is likely to produce artifacts, some of which may be hard to recognize in the absence of other controls. While in most cases FP fusions do not interfere with normal protein function, whenever possible, FP fusion proteins should be validated by immunostaining the corresponding endogenous protein in non-transfected cells and verifying similar patterns of localization.
FPs as expression reporters:
FPs are highly useful as quantitative expression reporters. By driving the expression of an FP gene by a specific promoter of interest, it is possible to produce an optical readout of promoter activity. Use of the brightest possible FP ensures the best dynamic range for such an experiment. Because dynamic localization is not generally an issue for expression reporter applications, it is possible to use non-monomeric FPs for this purpose, opening up additional possibilities for multiple wavelength imaging. In order to obtain more reliable quantitative data and to correct for likely variations between individual cells in expression reporter experiments, the use of two spectrally distinct (e.g. green and red) FPs is advisable. By driving expression of one FP with a constitutive promoter and a second FP with the promoter of interest, the ratio of the two signals provides a quantitative readout of relative activity. Averaged over many cells, this technique should provide statistical power necessary for quality expression level experiments. Because FPs normally have a very slow turnover rate in mammalian cells, it may be desirable to add a degradation tag to your FP to enhance temporal resolution when measuring highly dynamic promoter activity.
New Product of the Week 03-15-10 to 03-21-10: Oct4-Sox2 2-in-1 lentivirus ABP-SC-LVI2in1 for effective iPS generation link: http://www.allelebiotech.com/shopcart/index.php?c=132&sc=122.
Promotion of the Week 03-15-10 to 03-21-10: 5% off plate oligos at all scales! www.allelebiotech.com/allele3/Oligo_96Plate.php We are doing our “window promotion” again, during a hour-long window, get any Allele’s High efficiency competent cells at 30% regular price, the time will be announced tomorrow on our Facebook page.
Wednesday, December 16, 2009
mTFP1 is an excellent FRET donor
In one recent publication, Padilla-Parra et al (2) tested a number of different FRET couples to determine which was the best for fluorescence lifetime imaging (FLIM)-FRET experiments, and found that the mTFP1-EYFP pair was by far the best pair for FLIM-FRET. This group also confirmed that the fluorescence lifetime decay of mTFP1 fits well to a single exponential, and that the time constant for this decay is unaffected by photobleaching, making mTFP1 an excellent choice for any kind of fluorescence lifetime imaging applications, including FLIM-FRET. This group also notes that it is likely that the use of Venus or mCitrine variants in place of EYFP would improve the performance of this FRET pair even further.
In a mathematical analysis of the potential FRET efficiency of mTFP1 with Venus YFP, Day et al. (3) showed that compared with Cerulean (currently the brightest cyan Aequorea GFP variant), one can expect up to 17% better FRET efficiency using mTFP1. This group went on to characterize the mTFP1-Venus pair in live-cell FRET and FLIM-FRET experiments and showed that it worked as predicted in both cases. They also note that mTFP1 has superior brightness and photostability when compared to Cerulean in live cells, which is consistent with all in vitro data reported previously (1). In a related paper, Sun et al. (4) demonstrated that mTFP1 is also an excellent FRET donor for the orange fluorescent protein mKO2.
Together, these recent independent studies confirm that mTFP1 among the best options when choosing a fluorescent protein as a FRET donor. With its proven track record of successful fusions, mTFP1 is also an excellent all-around performer that will enhance almost any live-cell imaging experiment.
(1) Ai et al., (2006) Biochem. J. 400:531-540.
(2) Padilla-Parra et al., (2009) Biophys J. 97(8):2368-76.
(3) Day et al., (2008) J Biomed Opt. 13(3):031203.
(4) Sun et al., (2009) J Biomed Opt. 14(5):054009.
AlleleBlog Admin, by Nathan Shaner
Video of the month (NEW!): Protein Expression Systems on youtube (http://www.youtube.com/watch?v=n81orbUebsQ) and at our protein expression page.
Discount of the week (Dec 14-20): 15% off Phoenix Retrovirus Expression System 2.0 (with selection medium provided)
New product(s) of the week: 48 fluorescent protein fusions on ready-to-infect virus that get into primary mammalian cells as subcellular markers (http://www.allelebiotech.com/shopcart/index.php?c=197&sc=34), 20 infections, only $249 for a limited introduction time.
Friday, November 13, 2009
Take advantage of the brightest GFP for studying gene expression regulations
For studying factors that regulate mammalian gene expression, enzymes such luciferase and lacZ are traditionally used as reporters when operationally linked to promoters and enhancers. Fluorescent proteins are becoming more and more popular for such applications as instruments for reading fluorescence emitted from treated cells are becoming more available. Using fluorescent proteins as reporter eliminates the need for performing enzyme reactions with assay substrate kits. More importantly, fluorescence readings can be taken at any time point on LIVE cells.
Allele Biotech introduces to the market a set of gene expression reporter constructs based on mWasabi and its cyan relative mTFP1, as the new product of the week of 11/09/09 to 11/15/09. Choosing different versions within this vector group, promoters, enhancers, or DNA binding protein binding sites can be easily inserted and their effects of gene expression compared to those of controls.
Promotion of the week: Lentiviral particles expressing commonly used human cytokines at one-time discount.
News article in AlleleNews (to be published Thursday): Using GFP-tag in Immunoprecipitation to study DNA repair pathway factors.
Wednesday, September 2, 2009
Allele Annouces New Products Based on Camelid Antibodies
Applications of GFP-Trap may include ChIP-CHIP, CLIP, co-IP, enzyme activity analysis (see Allele Biotech’s product group main page for sample publications). Other products such as anti-RFP and anti-GFP monoclonal antibodies that may be used after GFP-fusion precipitation are now also available from Allele Biotech. “VHH fragments have great potentials in both therapeutic and basic research,” said Allele’s CEO Dr. Jiwu Wang, “The agreement will significantly strengthen Allele Biotech's position in the antibody field”. Allele Biotech started with a grant from the NIH in 2000 to develop ways to display and select antibodies. It participated in a collaborative project on yeast display for selecting antibodies against cancer antigens in 2007 for the NCI. After acquiring Orbigen in 2008, Allele has thousands of antibodies in its product line.
Tuesday, August 25, 2009
Immunoprecipitation Tags
Immunoprecipitation is a process of isolating a protein as an antigen by using antibodies against it. It is a powerful tool for studying proteins in biological samples and, in case of Co-IP (meaning immunoprecipitation of complexes containing a known antigen), for analyzing protein-protein interactions. Similar technologies such as chromatin immunoprecipitation (ChIP), RNA immunoprecipitation (RIP), or crosslinked and iImmunoprecipitation of RNA-protein complexes (CLIP) aid analysis of protein-DNA or protein-RNA interactions.
The major obstacle for achieving effective immunoprecipitation is the difficulty of finding usable antibodies against a target of interest. A common practice is to use tags that are fused to the C- or N-terminus of the target protein, thereby any validated, commercially available antibody can be used for co-IP in different experimental systems. However, caution must be exercised against potential interference of biological functions from the added tags. In general, one should choose tags that have been tested in many situations and proven non-interfering; still, each biological system is different. Independent validation or supporting data should be used when interpreting results from tag-based co-IP.
Tags are often selected based on high quality and commercially available antibodies. Most commonly used tags include: FLAG, Myc, HA, V5, T7, and His, which are quite small in size and in theory less likely to interfere. GST and GFP are in between 20-30kDa, but they are well documented to form self-contained and stable structures independent of their fusion partners and proved to not interfere in many cases. GST can bind to glutathione beads directly, therefore a top choice for pulldown experiments. GFP or other FPs as tags have the advantages of being also a visualization module to follow the protein both inside cells and during pulldown. However, previously available anti-GFP antibodies, either polyclonal or monoclonal, are not comparable to those against other tags, thereby limiting the use of GFP as fusion tag in pulldown experiments.
GFP-Trap, a recent addition to anti-tag antibodies, is an E. coli expressed, single domain fragment derived from camelid heavy chain antibodies with much higher stability, specificity, and affinity, making GFP based pulldown quantitative. This recent advancement should make GFP in line to become the most suitable tags for many aforementioned precipitation experiments.
Tuesday, June 16, 2009
Allele Will Bring a New Family of Fluorescent Proteins to the Market
These proteins were discovered in Amphioxus, a type of small fish that can be found in beach sand, which is believed to be a very primitive cordate species. Compared to jellyfish and coral, from which virtually all of the currently used fluorescent proteins were isolated, Amphoixus are closer to mammalians and their proteins may find great application in human cells and other commonly used animal models. In addition, there are a large number of protein variants that can provide different spectra and other important physical properties such as photostability and photoconvertability.
Allele Biotech’s plan is to first introduce several new fluorescent proteins of different colors to the market as immediate alternatives for fluorescent protein customers. The next step is to continue to evolve and mature these proteins to create more advanced proteins with desired properties suitable for live animal imaging or more advanced applications such as PALM/STORM and SIM. Allele Biotech has on its team of fluorescent protein research staff and collaborators, some of the most highly regarded scientists. With these resources, Allele Biotech plans on committing to long-term development of truly user-friendly fluorescence imaging products.
These new class of fluorescent proteins will be integrated into Allele Biotech’s current products including: retro/lentiviral vectors, baculovirus and bacmam systems, as well as iPSC and RNAi constructs.